Model > Evaluate > Evaluate classification
Evaluate model performance for (binary) classification
Response variable
Select the outcome, or response, variable of interest. This should be a binary variable, either a factor or an integer with two value (i.e., 0 and 1).
Choose level
The level in the response variable that is considered a success. For example, a purchase or buyer is equal to “yes”.
Predictor
Select one or more variables that can be used to predict the chosen level in the response variable. This could be a variable, an RFM index, or predicted values from a model (e.g., from a logistic regression estimated using Model > Logistic regression (GLM) or a Neural Network estimated using Model > Neural Network).
# quantiles
The number of bins to create.
Margin & Cost
To use the Profit and ROME (Return on
Marketing Expenditures) charts, enter the Margin for each
sale and the estimated Cost per contact (e.g., mailing
costs or opportunity cost of email or text message). For example, if the
margin on a sale is $10 (excluding the contact cost) and the contact
cost is $1 enter 10 and 1 in the Margin and
Cost input windows.
Show results for
If a filter is active (e.g., set in the Data >
View tab) generate results for All data,
Training data, Test data, or Both
training and test data. If no filter is active calculations are applied
to all data.
Plots
Generate Lift, Gains, Profit, and/or ROME charts.
Report > Rmd
Add code to
Report
> Rmd to (re)create the analysis by clicking the
icon on the bottom
left of your screen or by pressing ALT-enter on your
keyboard.
If we create a set of four charts in the Plots tab we can
add a title above the group of plots and impose a two-column layout
using patchwork as follows:
plot(result, plots = c("lift", "gains", "profit", "rome"), custom = TRUE) %>%
wrap_plots(plot_list, ncol = 2) + plot_annotation(title = "Model evaluation")The single plot can be customized using ggplot2 commands
(see example below)). See
Data
> Visualize for details.
Confusion matrix
Predicted probabilities probabilities selected through
Predictor are first converted to a class (e.g., a positive
or negative outcome) using the values entered in Margin and
Cost. It will be profitable to contact a customer if the
predicted probability of response exceeds Cost / Margin.
For example, if the break-even response rate is 0.1 and the predicted
probability of response is 0.25 that customer will be assigned the label
Positive. If, on the other hand, the predicted probability does
not exceed the break-even response rate a customer will be assigned a
Negative label.
Once each prediction has been converted to a class label (i.e., Positive or Negative) the result is compared to the values of the response variable. The following key measures are shown in the generated table for each predictor.
| Label | Description |
|---|---|
| TP (True Positive) | Number of cases where the positive prediction matches the positive outcome in the data |
| FP (False Positive) | Number of cases with a positive prediction but a negative outcome in the data |
| TN (True Negative) | Number of cases where the negative prediction matches the negative outcome in the data |
| FN (False Negative) | Number of cases with a negative prediction but a positive outcome in the data |
| total | Total number of cases (i.e., TP + FP + TN + FN) |
| TPR (True Positive Rate) | Proportion of positive outcomes in the data that received a positive prediction (i.e., TP / (TP + FN)). Also known as sensitivity or recall |
| TNR (True Negative Rate) | Proportion of negative outcomes in the data that received a negative prediction (i.e., TN / (TN + FP)). Also known as specificity |
| precision | Proportion of positive predictions with a positive outcome in the data (i.e., TP / (TP + FP)) |
| F-score | The harmonic mean of precision and TPR (sensitivity) |
| accuracy | Proportion of all outcomes that was correctly predicted as either positive or negative (i.e., (TP + TN) / total) |
| kappa | Corrects the accuracy measure for the probability of generating a correct prediction purely by chance |
| profit | Total profitability achieved by targeting all customers with a predicted probability above the break-even response rate |
| index | Index of relative profitability achieved across the selected
Predictor variables (maximum is 1) |
| ROME | Return on Marketing Expenditures (ROME) achieved by targeting all customers with a predicted probability above the break-even response rate |
| contact | Proportion of customers to contact, i.e., (TP + FP) / total |
| AUC | Area Under the ROC Curve (AUC). ROC stands for Receiver Operating Characteristic. |
Report > Rmd (confusion)
Add code to
Report
> Rmd to (re)create the analysis by clicking the
icon on the bottom
left of your screen or by pressing ALT-enter on your
keyboard.
Only kappa, index, ROME, and
AUC are plotted by default. It is possible to customize the
plotted results through Report > Rmd. To change the plot
use, for example:
The plot can be further customized using ggplot2
commands (see example below)). See
Data
> Visualize for details.
Download options
To download a table as a csv-files click the download button on the top-right of your screen. To download plots as png files click the download icon on the middle-right of your screen.
Example
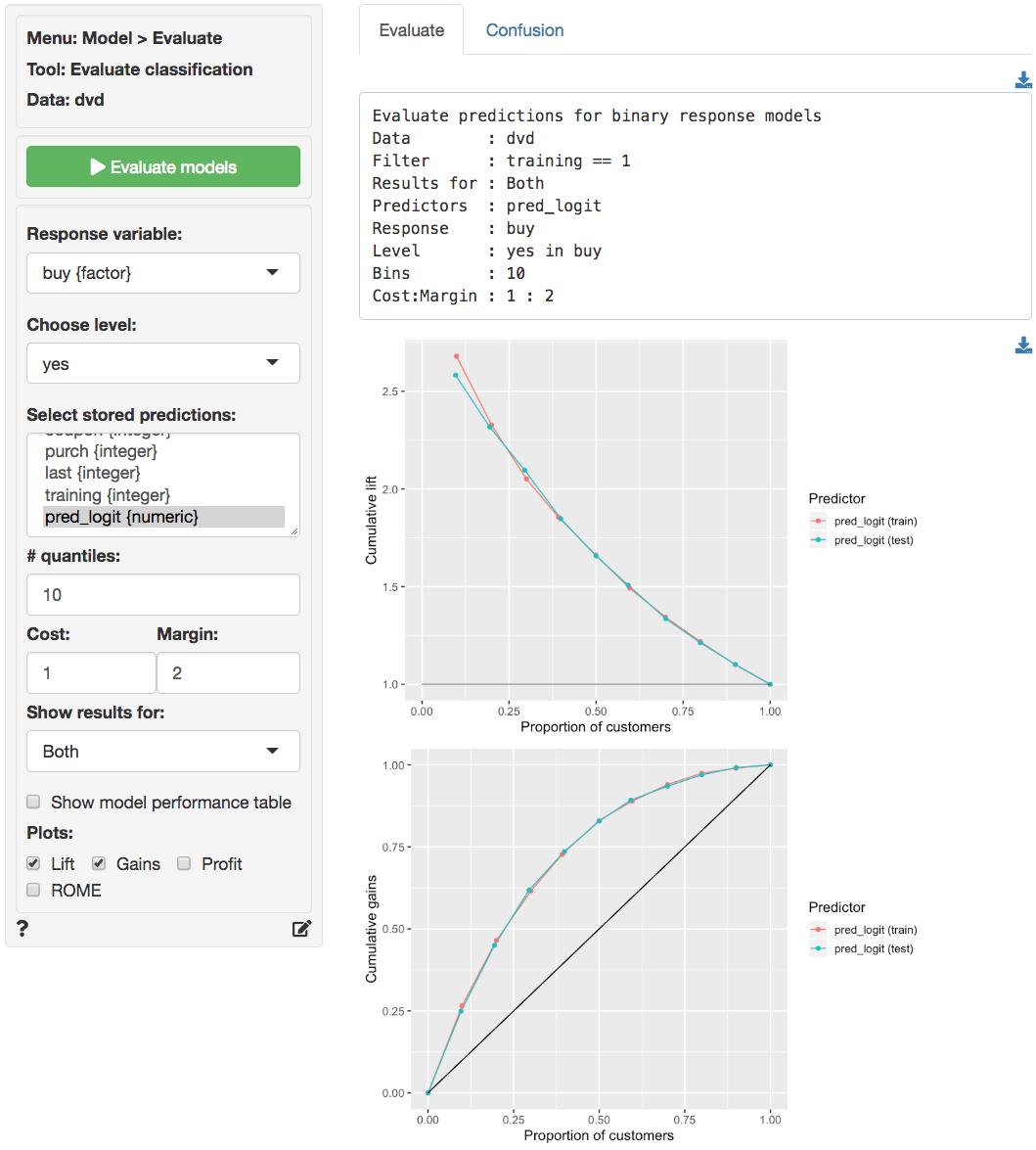
The Lift and Gains charts below show little evidence of overfitting and suggest that targeting approximately 65% of customers would maximize profits.
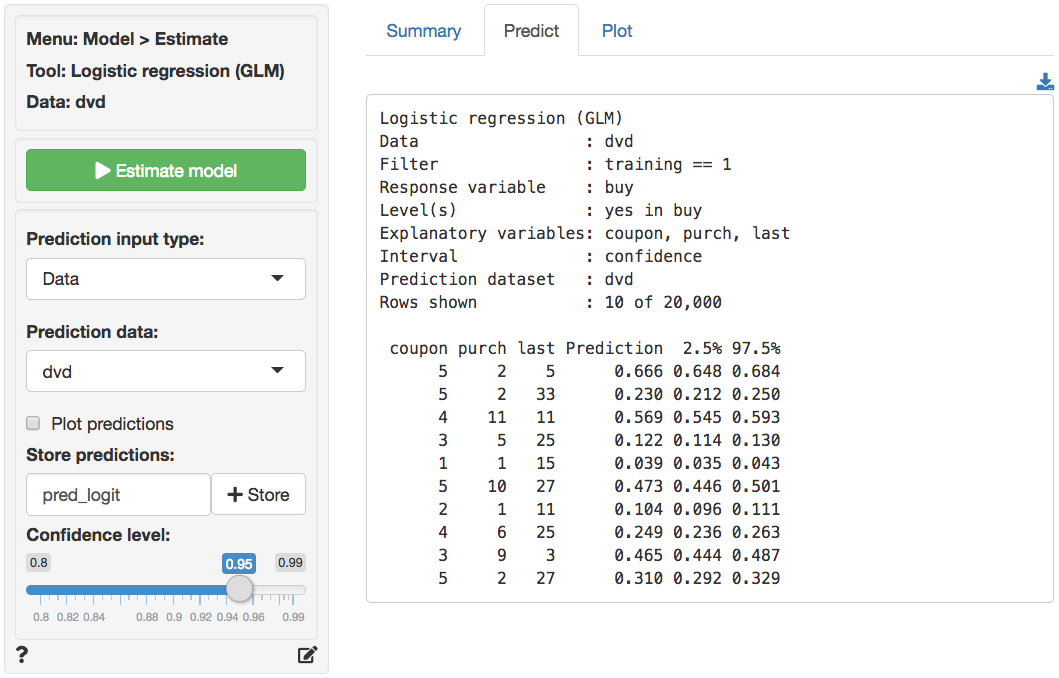
The prediction used in the screen shots above was derived from a
logistic regression on the dvd data. The data is available
through the Data > Manage tab (i.e., choose
Examples from the Load data of type drop-down
and press Load). The model was estimated using Model
> Logistic regression (GLM). The predictions shown below were
generated in the Predict tab.
R-functions
For an overview of related R-functions used by Radiant to evaluate (binary) classification models see Model > Evaluate classification